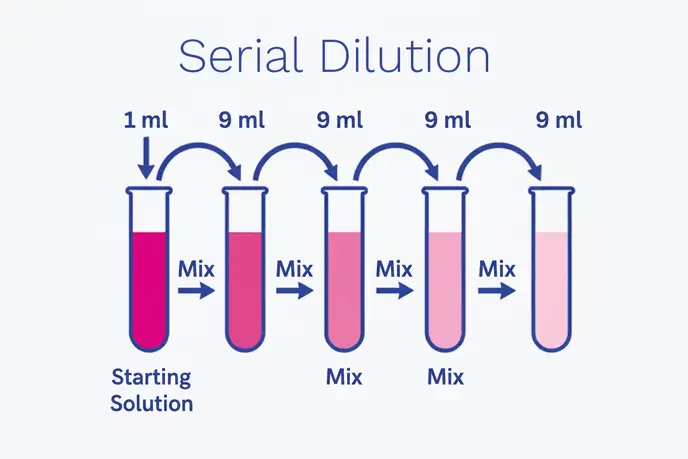
The arithmetic: Dilution factors, series, and log thinking
Serial dilutions are often expressed as Dilution Factors (DF). A single dilution step might be 1:10, 1:100 or 1:2, depending on the method and expected load. In a classic 10 fold series, each step reduces concentration by one log10. A 1:10 dilution repeated three times yields 1:1,000 (10-3).
Mathematically, the overall dilution factor is the product of each step’s factor. In a 2 fold serial dilution series (common in susceptibility testing or some titration formats), each step halves the concentration (2-n). In both cases, the principle is the same: controlled reduction so that measurement and interpretation become reliable.
The importance of ‘log thinking’ is not only microbiological - it also matters in assay interpretation. For example, in the Fluorescent Antibody Virus Neutralisation (FAVN) method (a cell based assay), the test uses a logarithmic scale for serial dilutions and then transforms results into a linear unit (IU/mL). A key consideration is that points higher on the log scale can translate into larger differences on the linear scale, making the reading less precise if the analyst is operating in the wrong region of the curve.
Hence, serial dilution is not just a mechanical activity - it is part of optimising where the laboratory seeks to measure on a response curve.
How serial dilution links to plate counts and bioburden trending
In routine bioburden testing - whether pour plate, spread plate or membrane filtration - the aim is to produce plates that are readable and representative. Most tests involved either adding a measured volume of sample to a Petri dish and pouring agar (such as 1 mL) or streaking/spreading a smaller aliquot (often 0.1 mL), then incubating.
When a sample is too concentrated, those approaches yield heavy growth, while when too dilute, the analyst may see zeros that fail to support confident conclusions (especially for low bioburden processes where the ‘absence’ of a specified or objectionable microorganism requires careful interpretation).
This is also why experienced teams treat dilution selection as a risk decision. If the analyst anticipates that a sample might be high (for example, from early-stage processing), they should build a dilution series that spans low to high. Conversely, for very clean samples (e.g., late-stage processing), the analyst might select a minimal dilution but ensure sufficient volume and replication to maintain sensitivity.
The ‘real world’ complication is that microbial populations are not always evenly dispersed - some bacteria clump, some adhere to particles, some survive better than others. As noted in internal material, colony counting becomes difficult with overcrowded plates, clumps, translucent colonies, edge effects or organisms that spread across agar - exactly the issues serial dilution is designed to prevent 3.
Technique matters: The most common failure modes
When investigations reference ‘laboratory error’, dilution errors tend to sit near the top of the list given that they are easy to make and hard to detect once the plate is incubated. Many out of specification checklists require a check to be made as to whether correct dilutions were undertaken, along with correct glassware and pipette volumes, as part of investigating laboratory error. This is a practical reminder that dilution is not merely arithmetic - it is execution.
The most common serial dilution failure modes include:
- Pipetting inaccuracies
Wrong volume, poor technique, uncalibrated pipette or using the wrong tip type
- Inadequate mixing
Failing to homogenise after each transfer leads to non representative aliquots (particularly with clumps or viscous samples)
- Carry-over contamination
Touching the pipette tip to non sterile surfaces, reusing tips where not permitted or splashing
- Wrong diluent
Touching the pipette tip to non sterile surfaces, reusing tips where not permitted or splashing
- Arithmetic/label errors
Tube labels not matching the intended series, transcription errors or mis-recorded dilution steps (a data integrity risk)
- Method mismatch
Using a dilution series designed for one plating volume (e.g., 1 mL pour plate) and applying it to another (e.g., 0.1 mL spread plate) without recalculating
The reason these errors matter is straightforward: the result often feeds into batch disposition, trending limits and deviation decisions. A single incorrect dilution can convert a true low-level contamination into a falsely alarming ‘action level’ count or it can mask a genuine issue by diluting it beyond detection.
Serial dilution in endotoxin and inhibition management
Serial dilution is also a ‘problem solver’ for assays that can be inhibited by product matrices. Take endotoxin testing, a routine dilution is normally adequate. However, if recovery/spike or assay response fails at that dilution, the analyst may need to go further out (e.g., from 1/80 to 1/100 to 1/200) while maintaining method validity.
This is not limited to endotoxin. Any assay with a curve (such as cell-based, chromogenic, turbidimetric, immunoassay tests) can have a usable range where precision is better. Many methods explicitly link serial dilution scale selection to precision, noting that operating at the wrong end of a log-scale curve increases inaccuracy. The broader lesson is such that serial dilution is part of assay design and interpretation, not merely sample preparation.
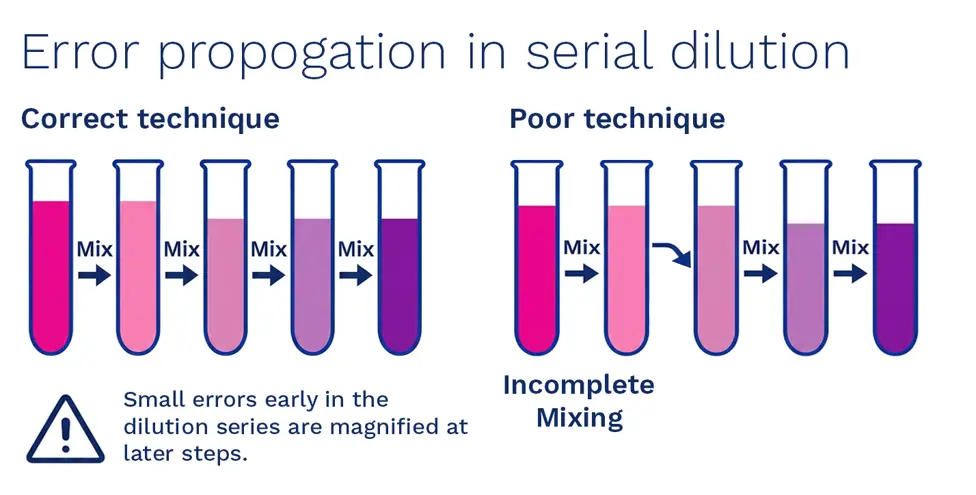
Image: Avoiding dilution errors (designed by Time Sandle)